Inferring time-derivatives including cell growth rates using Gaussian processes
The analysis code is written in Python 3.
If you are familiar with Python and uninterested in the GUI, then installing via pip is easiest:
pip install gaussianprocessderivatives
If not, all the prerequisites, including Python 3, can be installed at once using the Anaconda package. The software uses the
NumPy, SciPy, Matplotlib and Pandas modules, and installing Anaconda, or an equivalently package, is more straighforward than installing the modules individually, even if you already have Python on your computer.
You need three files: fitderiv.py,
fitderivgui.py, and gaussianprocess.py, but we have also included some example data and
an example script.
Download the software package (version 1.03)
These files must all be placed in the same
directory.
To get started, you should launch a terminal (a command prompt on Windows) and ensure that the
terminal accesses this directory before starting Python (using the cd command to change to the directory).
Contents
Running the GUI
Using a script
Trouble-shooting
Acknowledging the software
The GUI is written to provide a tool for estimating growth rates
from optical density data and uses that language (although it can
process any other
type of data too):
Start the GUI by typing
python fitderivgui.py
in a terminal (at the command prompt).
The GUI window will take a few seconds to appear
the first time it is called and should look like this on a Mac.
Clicking on Help will produce a new window with
detailed instructions.
Load the data by clicking on Load data. The data may
be a .xlsx, a .csv, or a .txt file. The first
column of data should be time and subsequent columns should be
measurements of optical density (OD). The data can also be arranged
in rows (i.e. the first row consists of time points, the second row
consists of OD measurements, etc.).
For a data set with four replicate measurements of
OD, the GUI window should now look something like this:
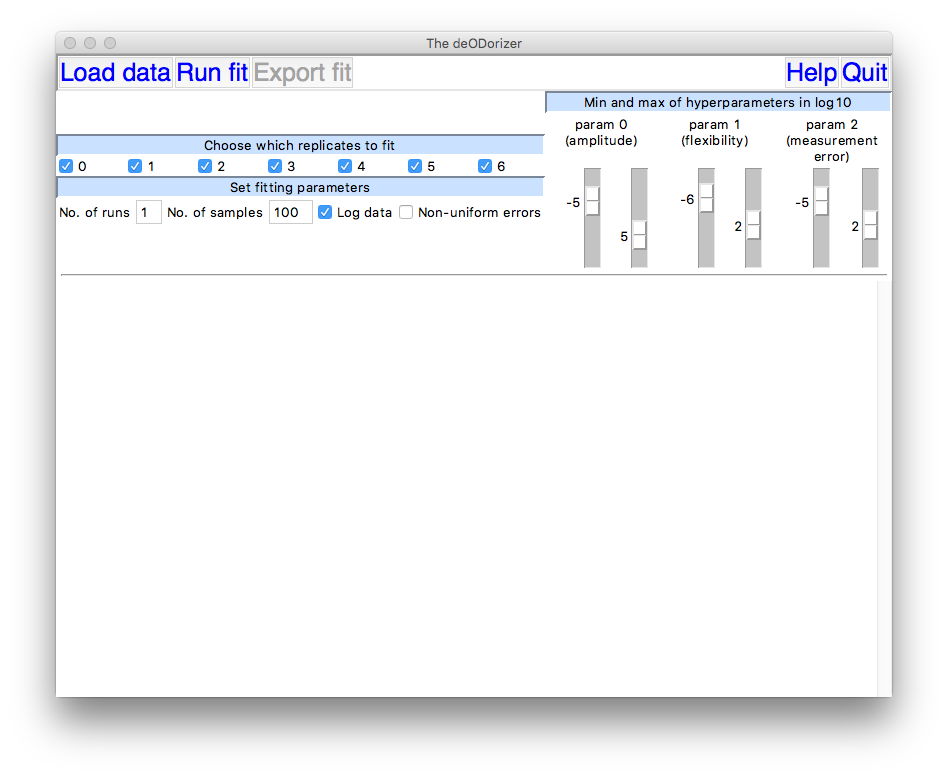
Run the fit by clicking on Run fit. If you wish you
can omit some replicates by de-selecting the appropriate box.
Once the fit has completed, a new window will display the results
graphically. The upper subplot shows the fit to the logarithm of the
OD data and the lower subplot shows the inferred growth rate. For
both, the dark blue line is the best-fit and the light blue is an
estimate of the error in this fit.
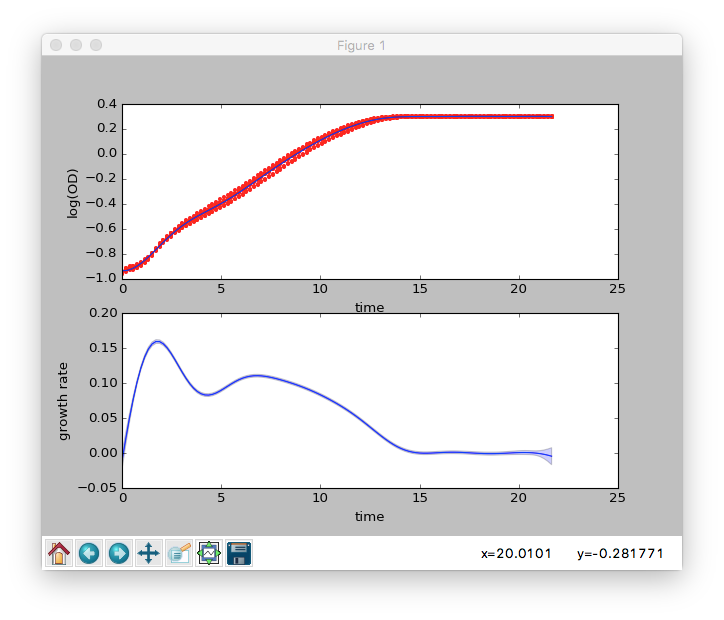
The main GUI window will display numerical results from the
fitting procedure including estimates for various summary statistics
of the growth curve:
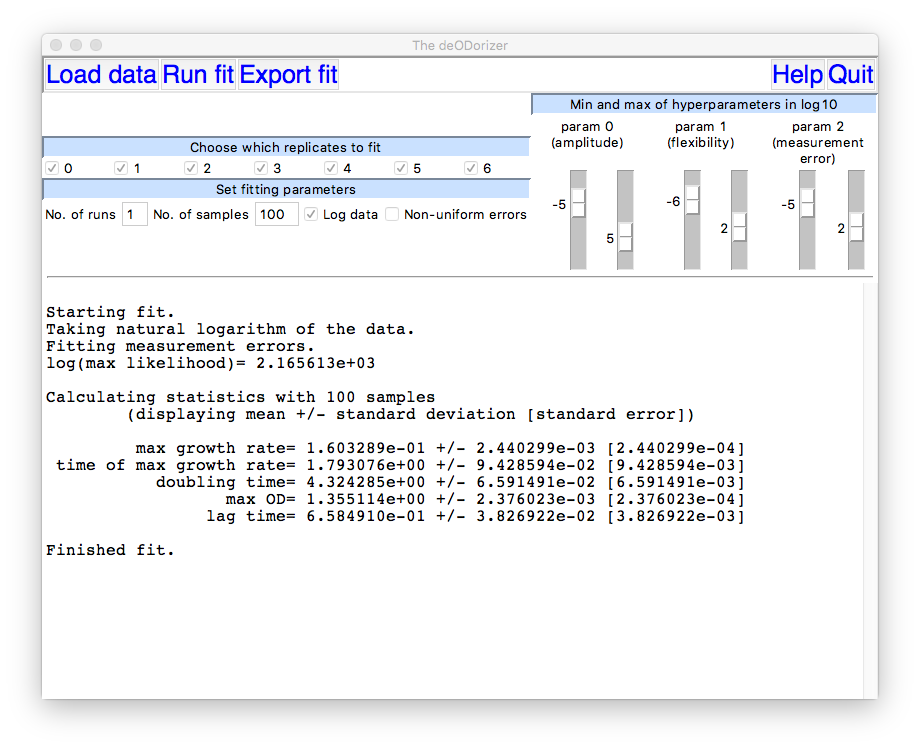
By sliding the
scale bars for the hyperparameters, you can change their upper and
lower bounds and then re-run the fit. The lower bound for the
hyperparameter for the amplitude (param
0) is here 10-5 and the upper bound is
105.
You may wish to change the bounds if one
hyperparameter is optimized to lie exactly on a bound (the GUI window will
display a warning) or if you dislike the smoothness of the best-fit
lines (change then the bounds for the flexibility parameter).
You can export the results to, for example, an .xlsx
file with Export fit. An additional file with the suffix
_stats.xlsx in this case will be created with the summary
statistics.
You can then either load a new data set with Load data
or shut down the entire GUI with Quit.
Further information is available under Help.
Figures 2A and 2B can be recreated in the GUI. For Figure 2B,
non-uniform errors should be toggled on.
We give below an example of Python code to call the algorithm using a
script where we do not rely on the default bounds for the
hyperparameters but specify particular values.
import numpy as np
from fitderiv import fitderiv
import matplotlib.pyplot as plt
# load data
d= np.loadtxt('data.txt')
t, od= d[:,0], d[:, 1:]
# redefine bounds and run inference
b= {0: [-1,5], 1: [-5,2], 2: [-7,1]}
q= fitderiv(t, od, bd= b)
# plot results
plt.figure()
plt.subplot(2,1,1)
q.plotfit('f')
plt.subplot(2,1,2)
q.plotfit('df')
plt.show()
# export results
q.export('results.xlsx')
Further information can be found by typing
help(fitderiv)
in for example IPython after the module has been imported.
If there is a poor fit to the data, try decreasing the upper bound on the hyperparameter for the measurement error.
If the inferred growth rate is too "noisy", try either increasing the lower bound on the hyperparameter for the measurement error to prevent over-fitting or decreasing the upper bound on the hyperparameter for the flexibility.
If the best-fit appears to change when you re-run the fitting, then the optimization is probably finding a local optimum. Try increasing the number of runs, say to either 5 or 10.
We tend to favour the Matern covariance function and, for data from our plate readers, typically raise the lower bound on the hyperparameter for the measurement error (the last one) to prevent overfitting.
Please cite:
PS Swain, K Stevenson, A Leary, LF Montano-Gutierrez, IBN Clark, J Vogel, and T Pilizota. Inferring time derivatives including growth rates using Gaussian processes Nat Commun 7 (2016) 13766



